A recent paper , Schwarz et al. 2010, was using kernel method to reconstruction the phylogenetic tree, which usually done by maximum likelihood estimation. Their using finite-state transducers(FST) to create a alignment-free kernel for evolutionary comparison of molecular sequence, and their call it a rational kernel approach. Their method overcome the gap in alignment sequence. As we known, the gap can influence the accuracy of phylogenetic tree.
Kernel method had approved to be a powerful tool for classification, and their method do help to classify the twilight-zone in very close sequence(see the following picture). The result in their paper is a new and accurate way of determining evolutionary distances in the twilight zone of sequence alignments that is suitable for large homologies datasets.
The method for phylogenetic/ phylogenomic reconstruction are still challenged problems in evolution biology. Schwarz et al. ‘s paper only do misclassification, maybe we can see the kernel method for estimating the divergence time, effective population size, recombination rate and mutation rate in nature population.
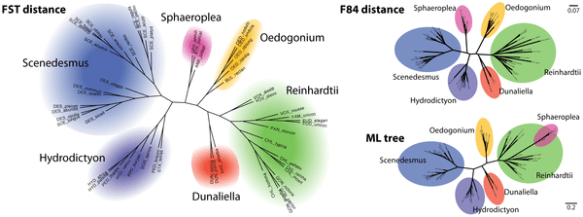
(A phylogenetic trees of the Chlorophyceae, which reconstructed by FST distance (left) using the full kernel score, F84 distance estimation on a Muscle alignment (top right) and maximum-likelihood tree on the same Muscle alignment (bottom right).)
Evolutionary Distances in the Twilight Zone—A Rational Kernel Approach
Roland F. Schwarz1*, William Fletcher2, Frank Förster3,Benjamin Merget3, Matthias Wolf3, Jörg Schultz3, Florian Markowetz1*
1 Cancer Research UK Cambridge Research Institute, University of Cambridge, Cambridge, United Kingdom, 2 Department of Genetics, Evolution and Environment and Centre for Mathematics and Physics in the Life Sciences and Experimental Biology, University College London, London, United Kingdom, 3Department of Bioinformatics, Biocenter, University of Würzburg, Würzburg, GermanyAbstract
Phylogenetic tree reconstruction is traditionally based on multiple sequence alignments (MSAs) and heavily depends on the validity of this information bottleneck. With increasing sequence divergence, the quality of MSAs decays quickly. Alignment-free methods, on the other hand, are based on abstract string comparisons and avoid potential alignment problems. However, in general they are not biologically motivated and ignore our knowledge about the evolution of sequences. Thus, it is still a major open question how to define an evolutionary distance metric between divergent sequences that makes use of indel information and known substitution models without the need for a multiple alignment. Here we propose a new evolutionary distance metric to close this gap. It uses finite-state transducers to create a biologically motivated similarity score which models substitutions and indels, and does not depend on a multiple sequence alignment. The sequence similarity score is defined in analogy to pairwise alignments and additionally has the positive semi-definite property. We describe its derivation and show in simulation studies and real-world examples that it is more accurate in reconstructing phylogenies than competing methods. The result is a new and accurate way of determining evolutionary distances in and beyond the twilight zone of sequence alignments that is suitable for large datasets.