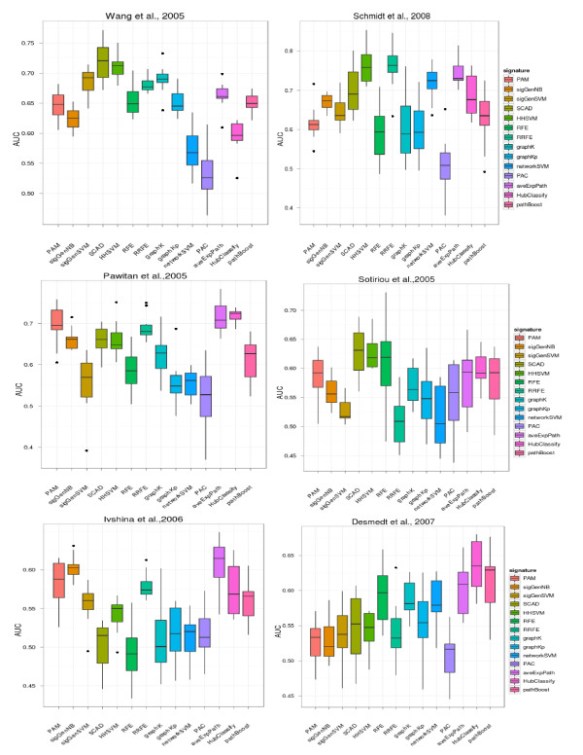
In our recent publication in BMC bioinformatics, we acompared a great deal of feature selection methods to finding prognostic biomakers in 6 breast cancer gene expresion data. No methods show significant performacne in prediction accuracy, feature selection stability and biogical interprety, which against previeous reseach results: current network-based appraoch did not show much benift in our analysis. Meanwhile, A group from NKI also show the simliar results in PloS One. The R codes for these algorithms in our paper is availiable as request.
Tag Archives: Machine Learning
Current approach in finding biomaker by means of mahcine learning
Biomarker Gene Signature Discovery Integrating Network Knowledge
Bonn-Aachen International Center for IT (B-IT), Dahlmannstr. 2, 53113 Bonn, GermanyAbstract: Discovery of prognostic and diagnostic biomarker gene signatures for diseases, such as cancer, is seen as a major step towards a better personalized medicine. During the last decade various methods, mainly coming from the machine learning or statistical domain, have been proposed for that purpose. However, one important obstacle for making gene signatures a standard tool in clinical diagnosis is the typical low reproducibility of these signatures combined with the difficulty to achieve a clear biological interpretation. For that purpose in the last years there has been a growing interest in approaches that try to integrate information from molecular interaction networks. Here we review the current state of research in this field by giving an overview about so-far proposed approaches.
When Machine learning meets molecular evolution
A recent paper , Schwarz et al. 2010, was using kernel method to reconstruction the phylogenetic tree, which usually done by maximum likelihood estimation. Their using finite-state transducers(FST) to create a alignment-free kernel for evolutionary comparison of molecular sequence, and their call it a rational kernel approach. Their method overcome the gap in alignment sequence. As we known, the gap can influence the accuracy of phylogenetic tree.
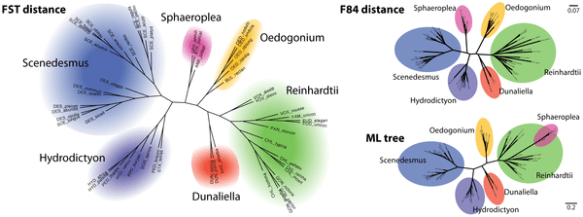
Kernel method had approved to be a powerful tool for classification, and their method do help to classify the twilight-zone in very close sequence(see the following picture). The result in their paper is a new and accurate way of determining evolutionary distances in the twilight zone of sequence alignments that is suitable for large homologies datasets.
The method for phylogenetic/ phylogenomic reconstruction are still challenged problems in evolution biology. Schwarz et al. ‘s paper only do misclassification, maybe we can see the kernel method for estimating the divergence time, effective population size, recombination rate and mutation rate in nature population.
(A phylogenetic trees of the Chlorophyceae, which reconstructed by FST distance (left) using the full kernel score, F84 distance estimation on a Muscle alignment (top right) and maximum-likelihood tree on the same Muscle alignment (bottom right).)
继续阅读