A recent paper , Schwarz et al. 2010, was using kernel method to reconstruction the phylogenetic tree, which usually done by maximum likelihood estimation. Their using finite-state transducers(FST) to create a alignment-free kernel for evolutionary comparison of molecular sequence, and their call it a rational kernel approach. Their method overcome the gap in alignment sequence. As we known, the gap can influence the accuracy of phylogenetic tree.
Kernel method had approved to be a powerful tool for classification, and their method do help to classify the twilight-zone in very close sequence(see the following picture). The result in their paper is a new and accurate way of determining evolutionary distances in the twilight zone of sequence alignments that is suitable for large homologies datasets.
The method for phylogenetic/ phylogenomic reconstruction are still challenged problems in evolution biology. Schwarz et al. ‘s paper only do misclassification, maybe we can see the kernel method for estimating the divergence time, effective population size, recombination rate and mutation rate in nature population.
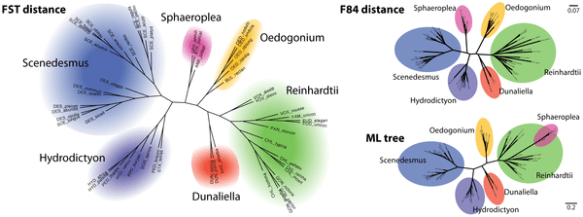
(A phylogenetic trees of the Chlorophyceae, which reconstructed by FST distance (left) using the full kernel score, F84 distance estimation on a Muscle alignment (top right) and maximum-likelihood tree on the same Muscle alignment (bottom right).)
继续阅读